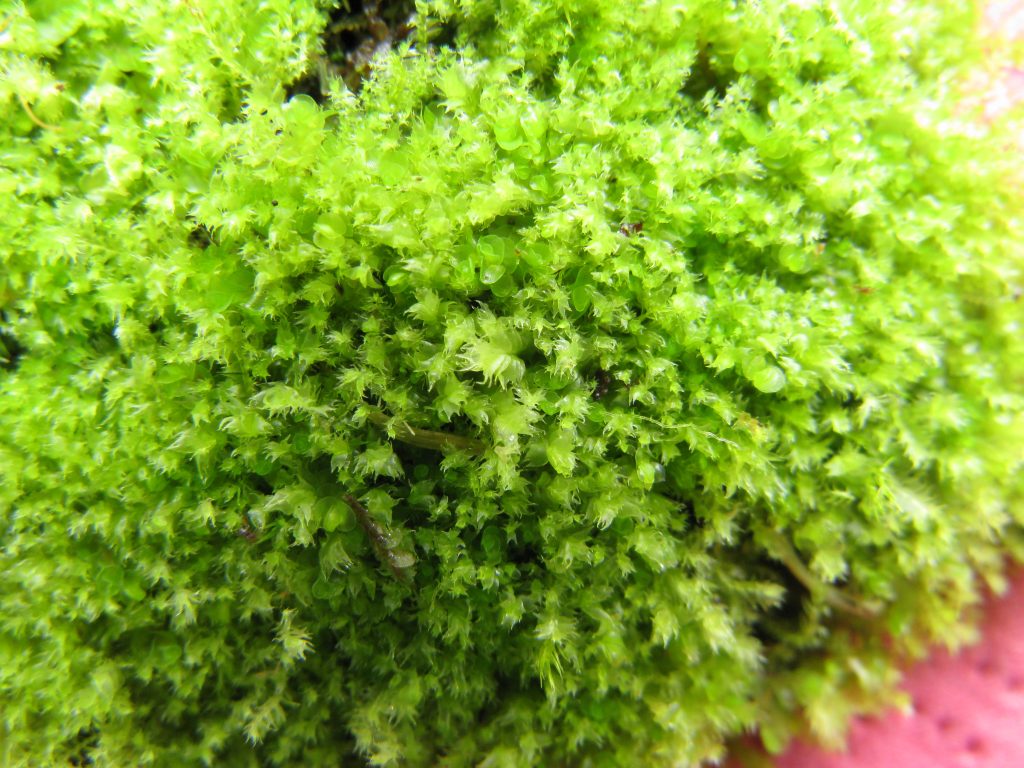
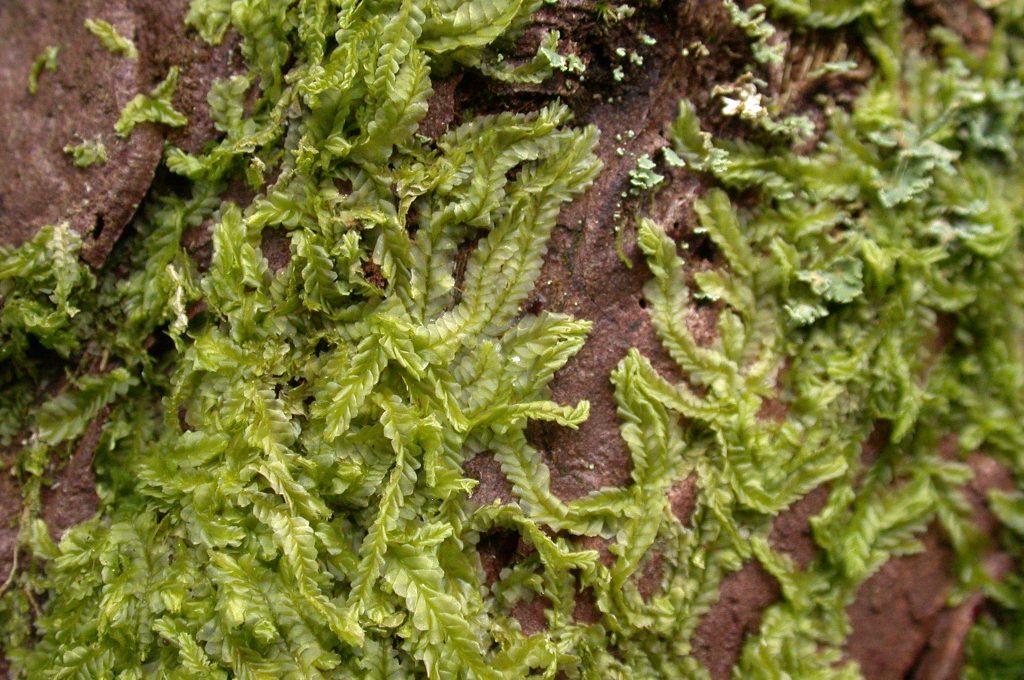
University of Edinburgh Biotechnology student Kenneth McKinlay’s 4th year honours project, 2013. Supervisors: Dr David Long, Dr Laura Forrest
Kenneth barcoded all six species of British Lophocolea, L. bidentata, L. bispinosa, L. brookwoodiana, L. fragrans, L. heterophylla and L. semiteres, attempting to get data from three plastid regions (rbcL, matK, psbA-trnH) and one nuclear region (ITS2). The data generated from the rbcL and psbA-trnH regions was effective in discriminating between all the species sampled; however useful data were not obtained from matK or ITS2.
Genetic markers:
1. rbcL: bidirectional sequence data was generated for 38 accessions.
2. matK: amplification was not successful with the primer sets used (LivF1A, LivR1A).
3. psbA-trnH: bidirectional sequence data was generated for 40 accessions.
4. ITS2: although PCR amplification was successful for 35 accessions, the low quality of many of the sequences generated, and the presence of clear heterozygous positions in sequence data from some accessions, made this data set problematic to analyse, so it was excluded from the study.
Species and trees:
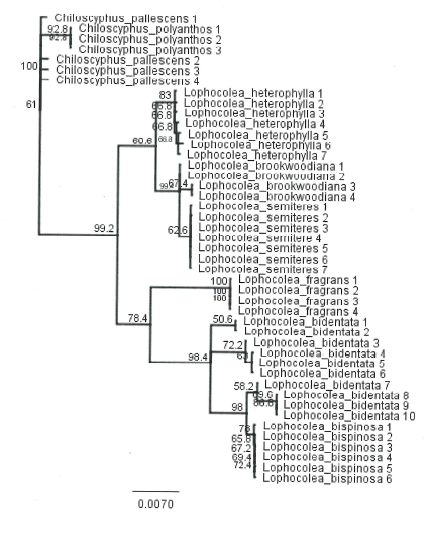
Distance tree generated using rbcL barcode sequence data for Lophocolea accessions, rooted on Chiloscyphus
L. fragrans – all accessions were genetically uniform, forming a monophyletic group.
L. heterophylla – although there was a little genetic variation, again, accessions of this species formed a distinct clade for both rbcL and psbA-trnH.
L. semiteres & L. brookwoodiana – these formed a single clade. All the accessions of L. semiteres (including material from the UK and Belgium) were genetically uniform, while two different genotypes were observed for L. brookwoodiana. While L. semiteres is known to be an introduced species in the UK, it’s possible that the three different genotypes in this clade represent separate introductions.
L. bispinosa – species formed a single genetically uniform group; this nests within a L. bidentata grade.
L. bidentata – accessions of this widespread and common species formed a grade, with three genetically distinct groups. One of these groups may represent L. cuspidata, a species that was sunk into L. bidentata by Bates and Walby in 1991, due to a lack of consistently distinguishing morphological characters. The results of this study suggest that a recircumscription of L. bidentata, “probably the commonest leafy liverwort in the British Isles” (Hodgetts, 2010), is required.
Related Posts
Student projects at RBGE: DNA barcoding British liverworts: Lophocolea
Student projects at RBGE: Barcoding British Liverworts: Plagiochila (Dumort.) Dumort.
Student projects at RBGE: Barcoding British Liverworts: Metzgeria



