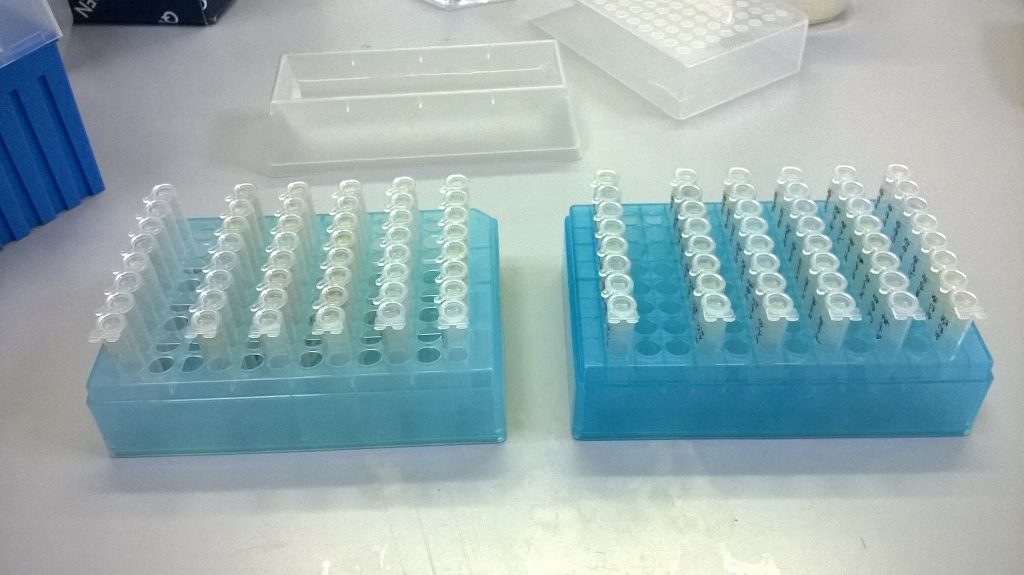
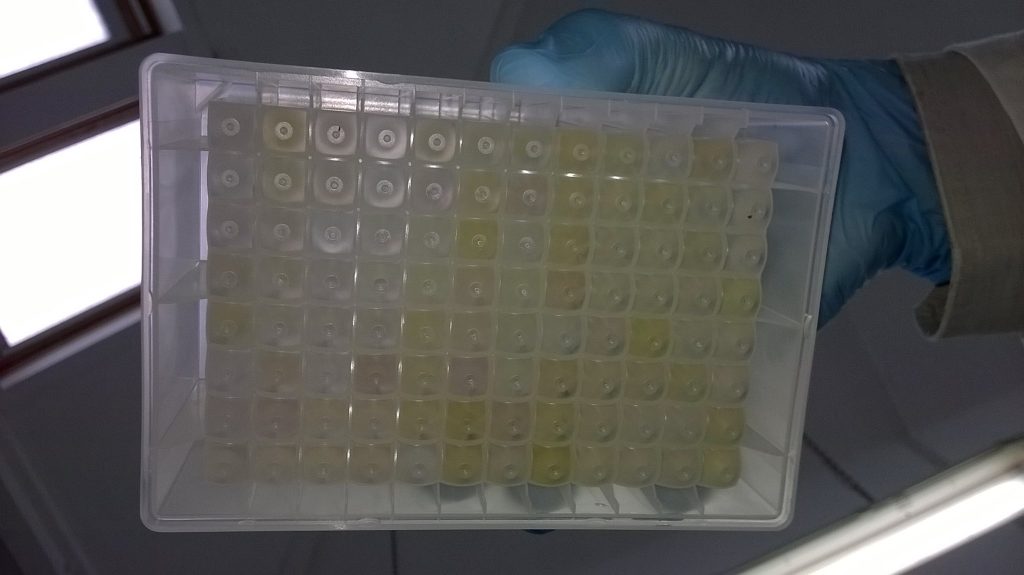
Just over a week into our current Synthesys-funded Schistidium project, and Wolfgang has picked through piles of packets of mosses, selecting the 96 that we would most like to get DNA sequences for, and putting tiny pieces of them into plastic tubes for DNA extraction.
Down in the basement, we have a storeroom with racks of lab coats, so we ventured down and found one that was a good fit for Wolfgang, to keep him from spilling chemicals down his regular clothes. We also got him some gloves; at RBGE, instead of latex, to which some people are allergic, we use nitrile. These serve two purposes – one, clearly, to protect our hands from any hazardous materials we might use, but in many cases in the molecular lab, the gloves are actually to protect our samples from us, so that we do not get our DNA or enzymes into the tubes we’re working with.
The first step of the extraction is the mechanical disruption of the plant tissue. Back to Biology 101, and a major difference between animal and plant cells is that plant cells have a tough cellulose-based cell wall around the outside of the cell membrane. To break through this cell wall, we add small tungsten beads to our tubes, and pop them into a TissueLysis machine which vibrates the tubes rapidly back and forth until the plants are rendered to a fine powder.
About half a millilitre (420 μl) of a salty, soapy buffer solution is added to this powder. Because we’re using tubes that are in strips of eight, we can use a multichannel pipette to dispense eight aliquots of the buffer at a time, reducing the amount of handling time. This first buffer helps break open the plant cell membranes, releasing the Schistidium DNA into solution. We put the tubes on a heated shaking block, leaving the samples for 1.5 hours at 65°C.
After the lysis step, the strips are centrifuged to remove bits of plant debris, 220 μl of the liquid Schistidium extraction is pipetted into a new deep 96-well plate, and 440 μl of a binding buffer is added, before 600 μl of the solution is transferred to yet another plate… This next plate contains a membrane to which the Schistidium DNA binds. While the DNA is stuck to the membrane, we pass four wash buffers over it, to remove compounds that might have co-extracted with the DNA, but that might inhibit some of our downstream reactions. These wash buffers contain quite high proportions of alcohol, so they don’t remove the DNA from the membrane (because DNA is not soluble in alcohol it doesn’t come out of precipitation). The final step in the process is to remove the DNA from the membrane in an elution step, using 100 μl of a water-based elution buffer per sample, in which the DNA can dissolve.
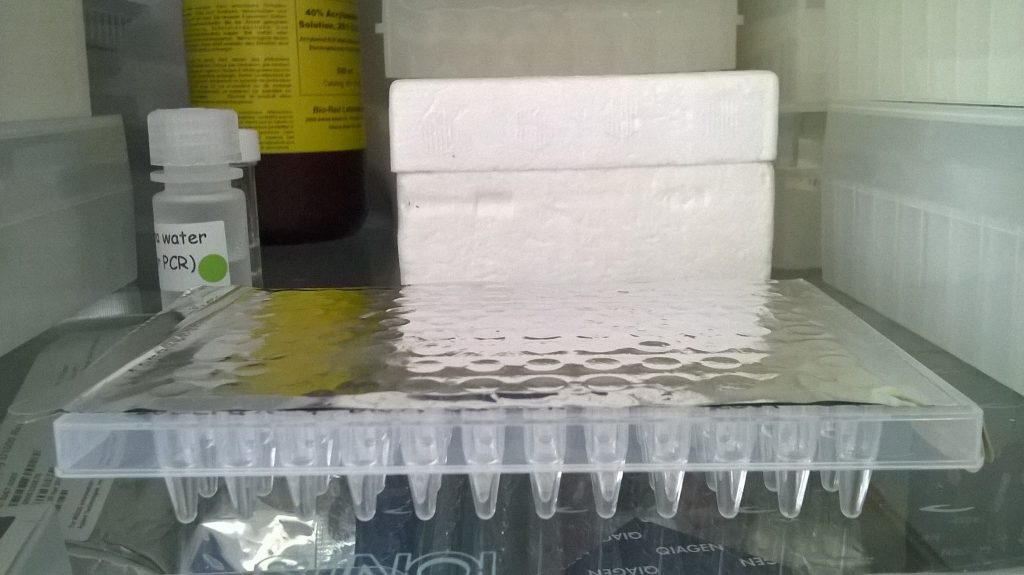
Once the 96 Schistidium DNA extractions are finished, the plate is sealed (with a thin sticky metal film), and left in the fridge (in this case, as we are going to use the DNA the next day), or in the freezer, until we are ready to move on to the next step.
Links to reports on Moss diversity in an artificial landscape, an EU Synthesys Access project with Dr Wolfgang Hofbauer at RBGE:
- Building on building mosses, a return to Schistidium in the built environment http://stories.rbge.org.uk/archives/24310
- Volunteering at the Botanics – bryophytes in our living landscape http://stories.rbge.org.uk/archives/24333
- Campylopus introflexus, an invasive alien on the glasshouse roof http://stor/ies.rbge.org.ukarchives/24359
- The trials and tribulations of a moss in the lab: DNA extraction http://stories.rbge.org.uk/archives/24399