It’s a horrible and unwelcome upheaval to have to change a protocol that works, but that’s the situation in which we have found ourselves with our semi-robotic DNA extraction system – the manufacturers upgraded the machine several months ago, and we can no longer get the kits that we have tried and tested on a huge range of plant and fungal material.
It’s not necessarily that the changes will make things worse, indeed they seem to add greater functionality to the system, but with two new kits to choose from, with different buffers and filters, which will better suit our needs? Things are always simpler when options are limited! The obvious thing is to run two similar plates of DNA extractions, one with each kit, and see if there’s a clear winner.
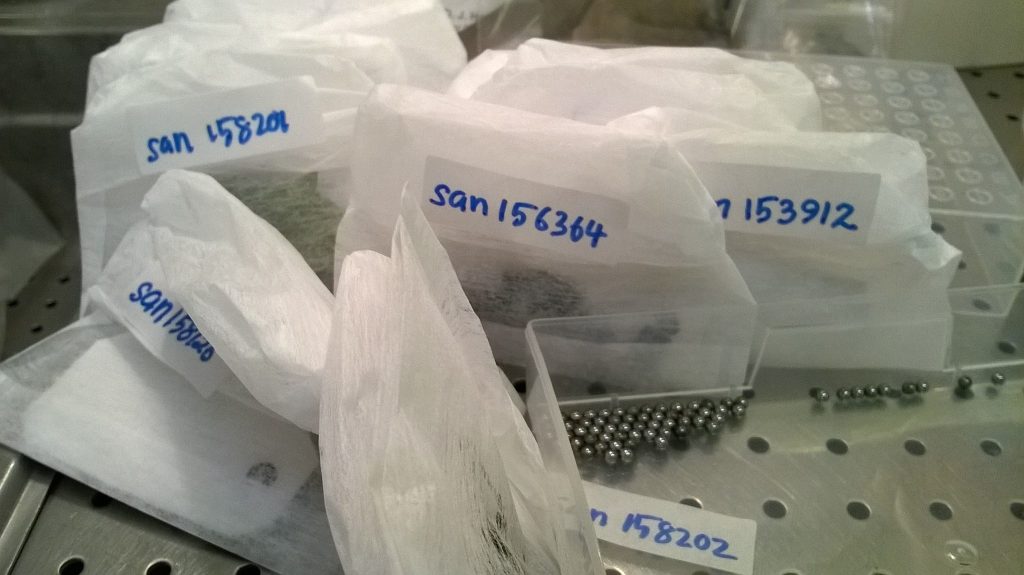
To do this, I have set up two matching plates of samples, each containing material from the same set of bamboos, conifers, gingers, lichens, liverworts, mosses and Sapotaceae samples. This covers a lot of the taxonomic range of things that we work on at RBGE; obviously we are looking for the protocol that works best across the widest range. I’ve also included a mixture of silica-dried plant material and herbarium specimens, as we’re involved in an EU-funded SYNTHESYS programme that investigates different ways of using our historic collections, including unlocking the molecular data stored within them. It will be useful to see if there’s much difference in how well the two kits deal with extracting the more degraded DNA that’s usually found in herbarium samples.
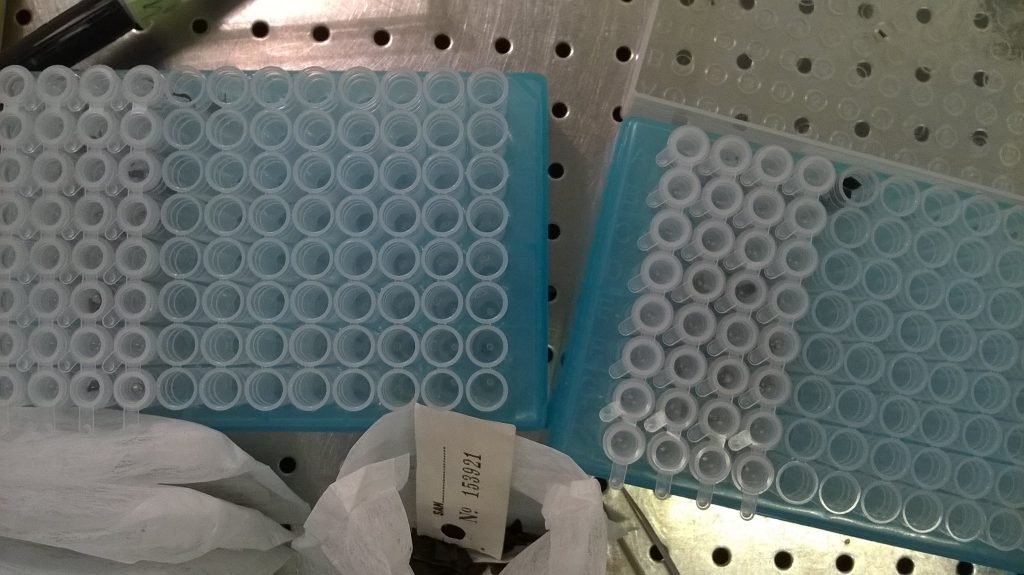
A small piece of each dried sample is placed in a small plastic tube, along with a couple of little round ball-bearings, which will be used to grind the sample into as fine a powder as possible. Carefully filling 96 tubes without dropping any bits into any of the adjacent tubes is fiddly and couldn’t really be described as fun, but it’s this sort of meticulous sampling work that is the foundation of any molecular genetic project.